Brhanu Fentaw Znabu
Ph.D. student in Biomedical Engineering at the University of Nebraska–Lincoln,
advised by Nicole R. Sexton at the
Nebraska Center for Virology
and co-advised by Qiuming Yao in the
School of Computing.
I build and evaluate genomic foundation models for biological sequence analysis.
My current work focuses on domain-adaptive pre-training of large language models (DNABERT-2) for
viral host-range prediction and epidemic emergence forecasting, with an emphasis on
rigorous leakage-aware evaluation and model interpretability.
I am also exploring discrete diffusion models and conditional generative frameworks
for protein variant design, combining protein language model representations with fitness-guided generation.

News
Publications
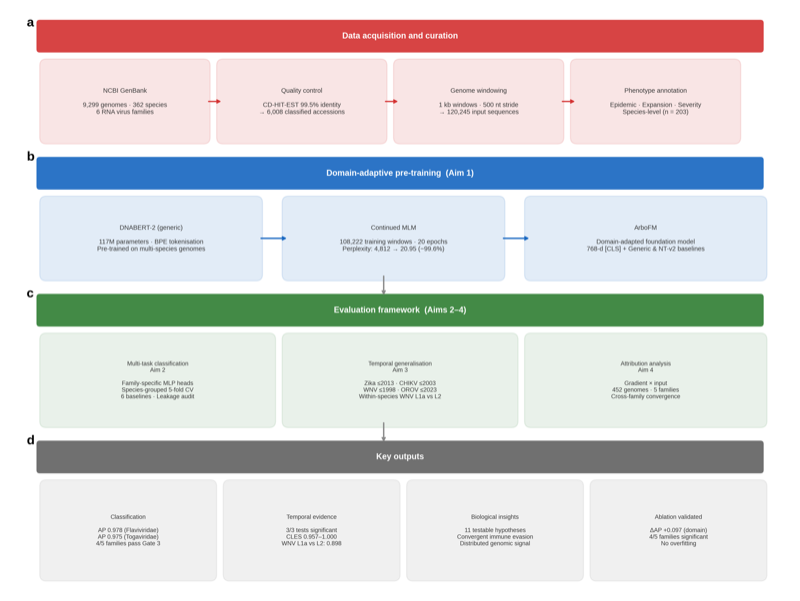
ArboFM: domain-adapted DNABERT-2 via continued MLM pre-training on 120K+ arbovirus genome windows (9,299 genomes, 362 species, 6 families). Predicts epidemic emergence with AP = 0.978 (Flaviviridae). Retrospective temporal validation detects Zika, chikungunya, and West Nile epidemic lineages before documented emergence.
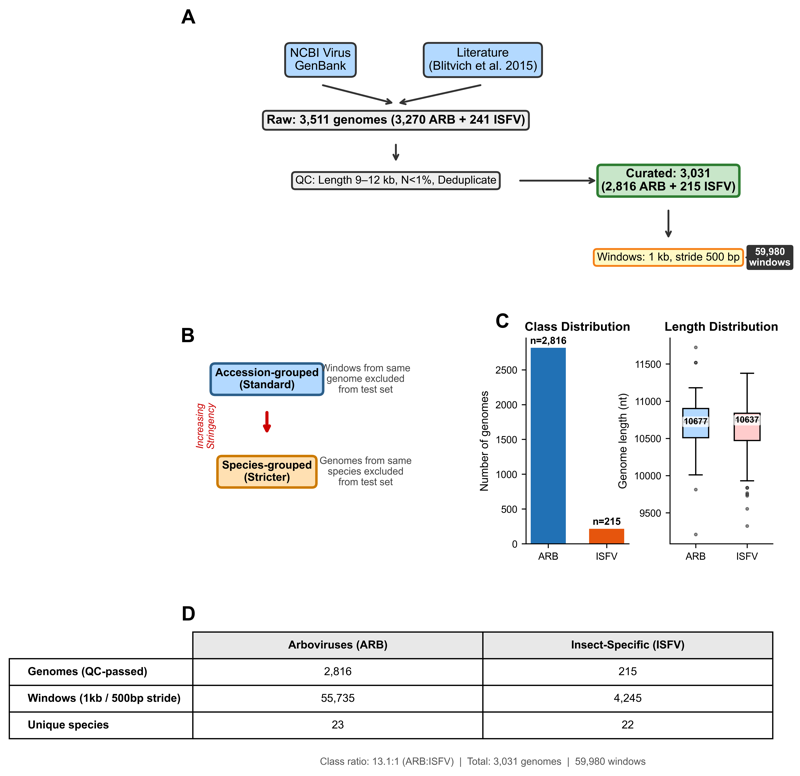
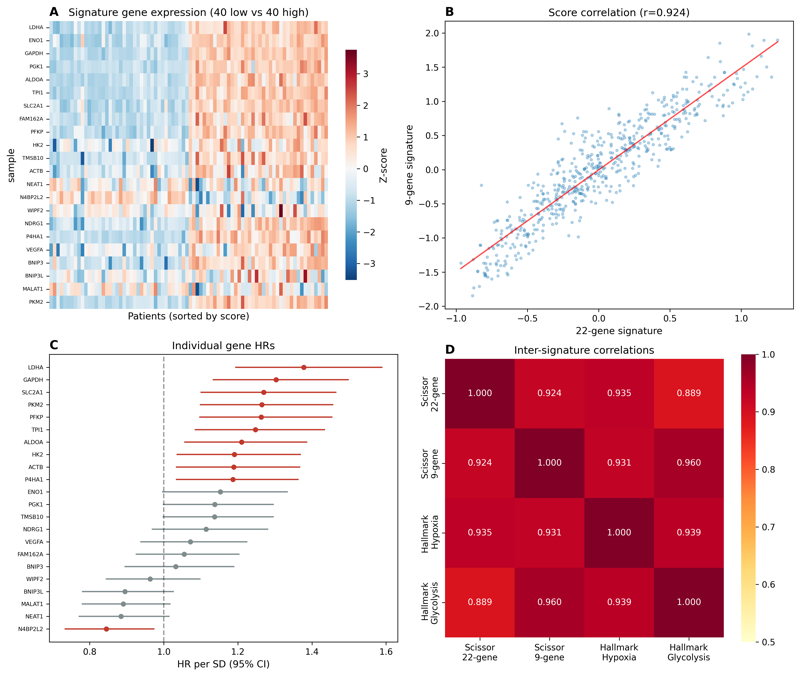
DNABERT-2 embeddings on 3,031 flavivirus genomes. Quantifies 15-percentage-point performance inflation from phylogenetic leakage. Identifies UpA-containing motifs linked to the OAS/RNaseL innate immunity pathway via genome-localized attribution.
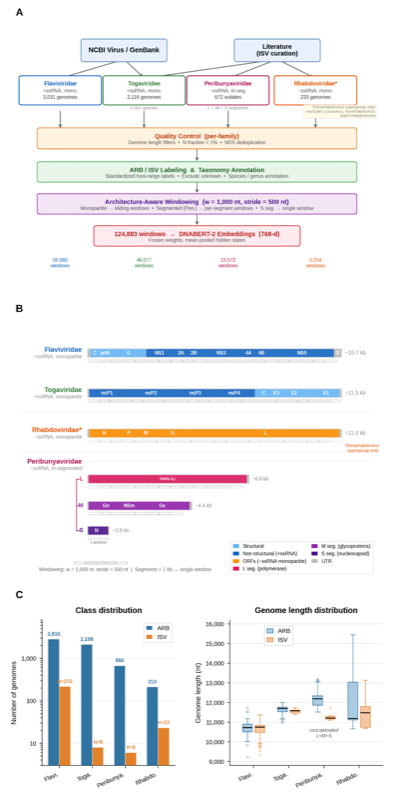
Cross-family transfer learning across 4 arbovirus families (6,052 genomes, 124,883 windows). Demonstrates host-switching grammar is predominantly family-specific (Jaccard ≈ 0), supporting convergent evolution.
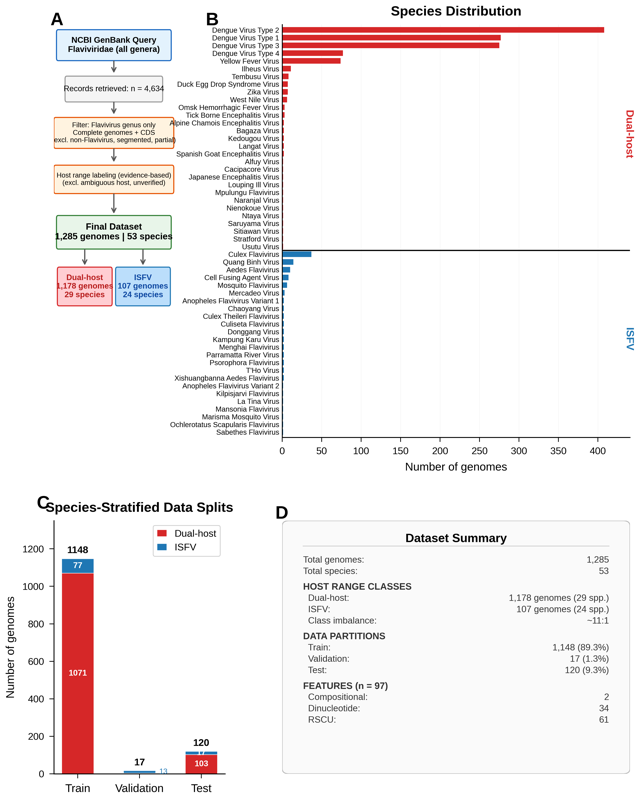
Species-stratified ML classifiers on 1,285 curated flavivirus genomes with 97 compositional features. Achieves PR-AUC = 1.000 for ISFV vs. dual-host classification. CpG suppression identified as the primary discriminative feature.
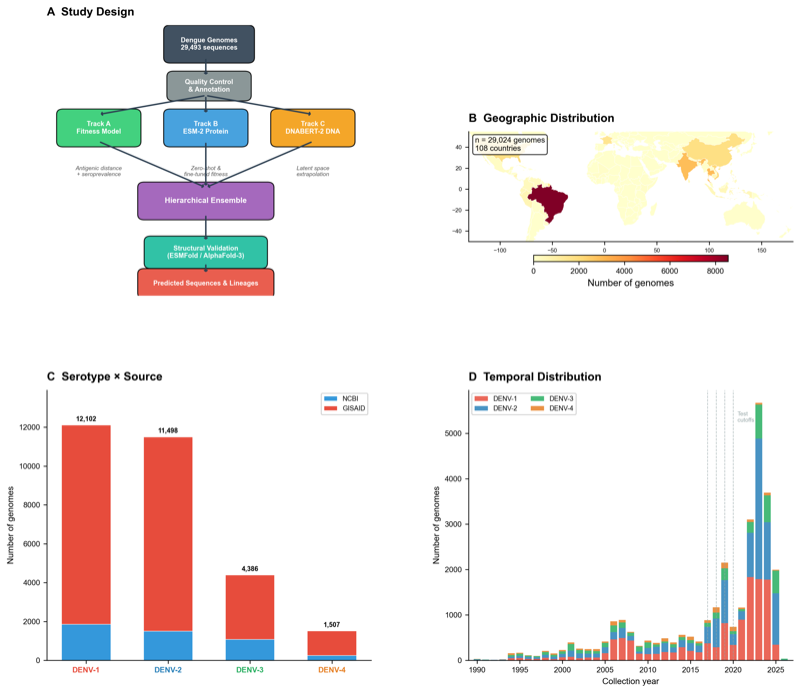
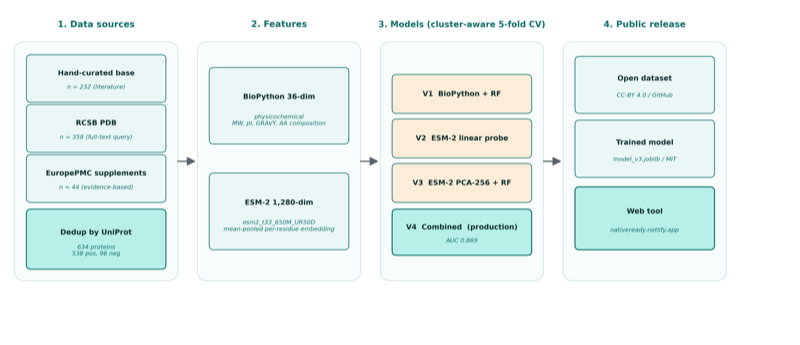
ViraPredict: multi-modal framework combining ESM-2 protein language models (650M params), fitness landscape modeling, and LoRA-fine-tuned DNABERT-2 with temporal embeddings on 29,493 curated dengue genomes across 130 countries.
Experience
About
My research sits at the intersection of machine learning and genomics. I develop foundation models and rigorous evaluation frameworks for biological sequence analysis, with applications in viral emergence prediction, host-range classification, and model interpretability. My work increasingly spans generative modeling, where I apply discrete diffusion and classifier-free guidance techniques to design protein variants with targeted functional properties. I work primarily with PyTorch, Hugging Face Transformers, and the SciPy/PyData ecosystem on HPC GPU clusters.
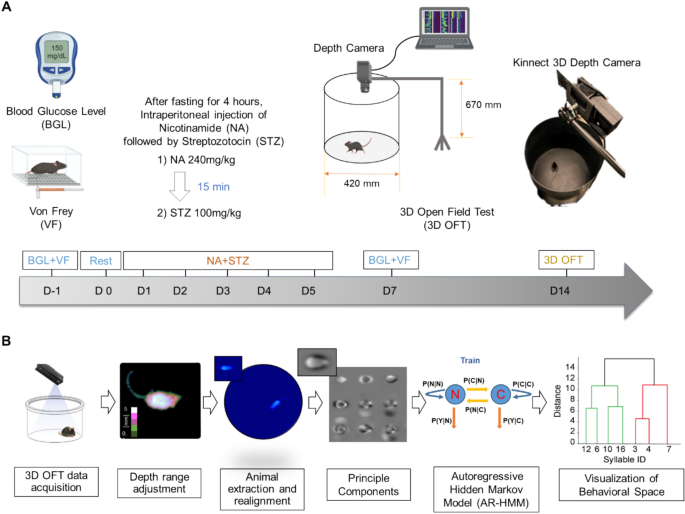
Prior to my PhD, I earned an M.Sc. in Biomedical Science and Engineering from Gwangju Institute of Science and Technology (GIST), South Korea, where I applied unsupervised machine learning (autoregressive models, Hidden Markov models) to 3D behavioral phenotyping in diabetic neuropathy mouse models. I also hold a B.Sc. in Biomedical Engineering from Jimma University, Ethiopia, graduating in the top 2% of my Biomedical Engineering class.
When I'm not at the computer, you'll find me exploring coffee shops ☕ or at the gym 🏋.